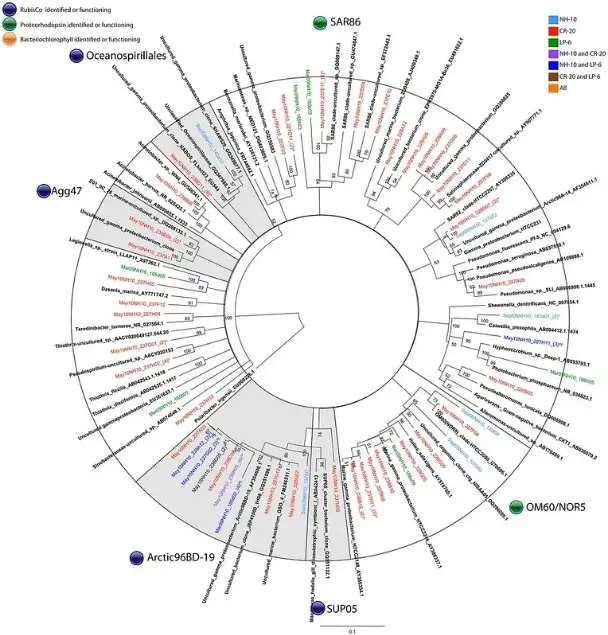
Phylogenetic trees are essential tools in the study of evolutionary relationships among species, offering insights into how various organisms have diverged and evolved over time. These trees, graphical representations of evolutionary ancestry, serve as a window into the history of life on Earth, tracing the lineage of genes, species, or larger groups. As scientific understanding deepens, methodologies to construct these trees have evolved, with UPGMA (Unweighted Pair Group Method with Arithmetic Mean) and Neighbor Joining (NJ) being two prominent techniques.
The difference between UPGMA and Neighbor Joining lies primarily in their approach and assumptions. UPGMA is a simple, distance-based method that assumes a constant rate of evolution across all lineages, producing a rooted tree. In contrast, Neighbor Joining does not make this assumption, resulting in an unrooted tree that can be more accurate in depicting evolutionary distances when rates of evolution vary among lineages.
Understanding these methods is crucial for interpreting the evolutionary history they reveal. UPGMA, with its simplicity and speed, is well-suited for analyzing sequences with uniform rates of evolution. Meanwhile, Neighbor Joining offers flexibility and accuracy for more complex datasets where evolutionary rates may differ. These tools not only provide foundational knowledge for evolutionary biology but also have applications in various fields, including conservation genetics, epidemiology, and the study of biodiversity.
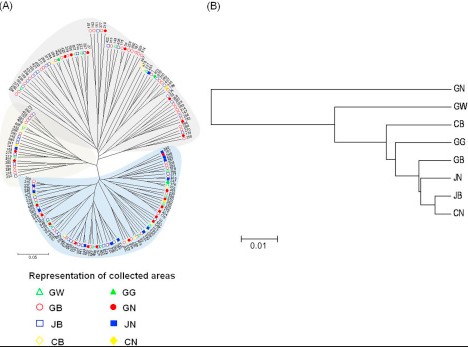
Phylogenetic Trees: A Primer
What Are Phylogenetic Trees?
Phylogenetic trees are visual representations of the evolutionary relationships among various biological species or entities based upon similarities and differences in their physical or genetic characteristics. These trees are constructed using a variety of algorithmic approaches to depict how species are related to each other in a branching diagram that reflects their evolutionary history. The root of the tree represents the common ancestor shared by all the branches, and each branch point denotes a divergence from this common lineage, highlighting the speciation events that lead to the formation of new species.
Purpose and Applications
Phylogenetic trees serve multiple purposes in biological research and application. Their primary use is in the study of evolutionary biology, where they help scientists understand the evolutionary processes that give rise to biodiversity. Beyond this, phylogenetic trees are crucial in fields such as conservation biology, where they can guide decisions on species preservation efforts by identifying species with few close relatives. In medicine, they are used to trace the transmission paths of infectious diseases and in the development of vaccines. Additionally, phylogenetic analysis plays a vital role in agriculture for crop improvement and in understanding the evolution of pests and diseases.
UPGMA Overview
Definition and Process
The Unweighted Pair Group Method with Arithmetic Mean (UPGMA) is a simple agglomerative (hierarchical) clustering method used in bioinformatics for constructing phylogenetic trees. The process involves the following steps:
- Calculating the distance matrix: Determining the pairwise distance between all taxa (species or sequences) based on their genetic or phenotypic characteristics.
- Cluster formation: Identifying the pair of taxa with the shortest distance and grouping them into a single cluster.
- Updating the distance matrix: Calculating the distance of this new cluster to all other taxa or clusters using the arithmetic mean of distances.
- Repeating the process until all taxa are clustered into a single phylogenetic tree.
Key Characteristics
UPGMA is known for its simplicity and speed, making it suitable for preliminary analyses or when working with small datasets. The method assumes a constant rate of evolution across all branches of the tree (molecular clock hypothesis), which means it is best used for groups believed to evolve at similar rates.
Advantages
- Ease of implementation and interpretation.
- Fast computation, ideal for small datasets.
- Provides a clear and simple representation of evolutionary relationships, especially when the molecular clock assumption holds true.
Neighbor Joining Overview
Definition and Process
Neighbor Joining (NJ) is a distance-based method used to construct phylogenetic trees. Unlike UPGMA, NJ does not assume a constant rate of evolution across all lineages. The NJ process is characterized by:
- Initial distance matrix calculation: Similar to UPGMA, but with a focus on minimizing the total branch length.
- Selecting pair of taxa: Choosing two taxa that, when joined, have the least effect on the overall length of the tree.
- Forming new nodes: Creating a new node representing the joined taxa and recalculating distances between this new node and all other taxa or nodes.
- Iterative joining: Repeating the process until all taxa are incorporated into the tree.
Key Characteristics
Neighbor Joining is valued for its ability to handle datasets where evolutionary rates vary among lineages. It is more flexible than UPGMA and often provides a more accurate representation of the evolutionary history of the taxa being analyzed.
Advantages
- No assumption of a constant rate of evolution, making it more broadly applicable.
- Efficiently handles large datasets, providing quicker results than many other methods.
- Often produces trees that are closer to the true evolutionary history of the taxa.
Comparing UPGMA and NJ
Algorithmic Differences
The fundamental distinction between UPGMA and NJ lies in their algorithmic approaches to tree construction. UPGMA uses a simple hierarchical clustering method, grouping taxa based on their average distances. In contrast, Neighbor Joining optimizes tree topology based on the minimization of total branch length, allowing for variable rates of evolution across branches.
Assumptions and Implications
Assumptions in UPGMA
- Constant rate of evolution (molecular clock) across all lineages.
- Evolutionary distances between taxa can be accurately represented by their pairwise distances.
Assumptions in NJ
- No molecular clock assumption, accommodating variable evolutionary rates.
- Focuses on minimizing the total tree length, aiming for an optimal tree structure.
Accuracy and Complexity
Accuracy in reflecting true evolutionary history
- NJ is generally considered more accurate than UPGMA, especially for complex datasets with unequal rates of evolution.
- UPGMA’s reliance on the molecular clock hypothesis can lead to misrepresentation of evolutionary relationships in the absence of a constant rate of evolution.
Computational complexity
- UPGMA is simpler and faster, making it suitable for small datasets or preliminary analysis.
- NJ, while more computationally intensive, is capable of handling larger datasets and providing more accurate results when the molecular clock assumption does not hold.
Use Cases and Preferences
When to Use UPGMA
UPGMA (Unweighted Pair Group Method with Arithmetic Mean) is a straightforward and efficient method for constructing phylogenetic trees, best suited for certain scenarios:
- Small datasets: Its simplicity and speed make it ideal for analyzing a limited number of sequences.
- Preliminary analysis: UPGMA can provide a quick overview, helping to identify major relationships before applying more complex methods.
- Uniform evolutionary rates: It works well when the assumption of a constant rate of evolution across all lineages (molecular clock) is reasonable.
- Educational purposes: Due to its simplicity, UPGMA serves as an excellent tool for teaching the principles of phylogenetic tree construction.
When to Use Neighbor Joining
Neighbor Joining (NJ) offers greater flexibility and accuracy in many scenarios, making it a preferred choice for:
- Large datasets: NJ is efficient and scalable, capable of handling larger numbers of sequences without significant loss in performance.
- Variable evolutionary rates: It does not assume a constant rate of evolution, making it suitable for datasets where evolutionary rates differ among lineages.
- Refined phylogenetic analysis: NJ often produces trees that are closer to the true evolutionary history, especially in complex evolutionary scenarios.
- Comparative genomics and evolutionary studies: Its ability to more accurately reflect evolutionary distances makes it valuable in these fields.
Limitations and Challenges
UPGMA Limitations
- Molecular clock dependency: The assumption of a constant rate of evolution is a significant limitation, as it is often not met in real-world data.
- Less accurate for complex data: UPGMA may not accurately depict the evolutionary relationships in datasets with variable rates of evolution.
- Over-simplification: While its simplicity is an advantage in some scenarios, it can also lead to oversimplification of the phylogenetic relationships.
NJ Limitations
- Computational intensity: For extremely large datasets, the computational resources required can be a limiting factor.
- Optimization issues: Finding the optimal tree can be more challenging due to the algorithm’s complexity, potentially leading to inaccuracies if not properly managed.
- User expertise: The interpretation of NJ trees, given their complexity, may require a higher level of expertise in phylogenetic analysis.
Recent Advances and Tools
Software for UPGMA
Recent years have seen the development of various software tools that implement the UPGMA algorithm, making phylogenetic analysis more accessible:
- MEGA: Offers a user-friendly interface for conducting phylogenetic analysis, including UPGMA, with tools for editing and visualizing trees.
- Phylip: A suite of programs for inferring phylogenies, which includes an implementation of UPGMA, suitable for both educational and research purposes.
Software for Neighbor Joining
Similarly, numerous tools now exist for performing Neighbor Joining analyses, facilitating more accurate phylogenetic trees:
- MEGA: Also supports Neighbor Joining, providing a comprehensive suite for phylogenetic analysis with various options for customization.
- SplitsTree: A powerful tool for analyzing and visualizing evolutionary relationships, SplitsTree offers advanced features for constructing Neighbor Joining trees and exploring the resulting data.
Innovations in Phylogenetic Analysis
The field of phylogenetic analysis continues to evolve, with ongoing research leading to new methodologies and enhancements to existing algorithms:
- Hybrid methods: Combining the strengths of different algorithms, such as UPGMA for initial clustering and NJ for refinement, to improve accuracy and efficiency.
- Machine learning approaches: Leveraging AI and machine learning to predict evolutionary relationships and optimize phylogenetic tree construction.
- Cloud computing: Utilizing cloud-based platforms to handle the computational demands of large-scale phylogenetic analyses, making advanced methods more accessible.
FAQs
What is a Phylogenetic Tree?
A phylogenetic tree is a diagram that represents the evolutionary relationships among various biological species or other entities based on similarities and differences in their physical or genetic characteristics. It illustrates how different species have branched out from common ancestors over time, providing insights into the history of life.
How Do UPGMA and Neighbor Joining Differ?
UPGMA and Neighbor Joining differ in their assumptions about evolutionary rates and the structure of the resulting tree. UPGMA assumes a constant rate of evolution across all branches, producing a rooted tree, whereas Neighbor Joining does not assume a constant rate, allowing it to generate an unrooted tree that may more accurately reflect the evolutionary distances among species when rates vary.
When Should You Use UPGMA?
UPGMA should be used when the data suggests a constant rate of evolution across all lineages. It is particularly effective for analyzing sequences from closely related species or within populations where evolutionary rates are expected to be uniform, making it a quick and simple method for constructing phylogenetic trees.
When Is Neighbor Joining Preferred?
Neighbor Joining is preferred when dealing with complex datasets where evolutionary rates may not be uniform across all lineages. It is flexible and more accurate in these scenarios, making it suitable for a wide range of evolutionary studies, including those involving highly divergent species or genes.
Conclusion
The choice between UPGMA and Neighbor Joining methods for constructing phylogenetic trees depends on the nature of the data and the specific requirements of the study. Both methods have their unique advantages and limitations, shaping how scientists understand evolutionary relationships. As the field of evolutionary biology progresses, these tools will continue to evolve, offering more refined insights into the tree of life.
Understanding the differences and applications of UPGMA and Neighbor Joining enhances our ability to interpret the complex web of life on Earth. The future of phylogenetic analysis promises further advancements, integrating new data and computational techniques to reveal even deeper insights into evolutionary histories and the connections that bind all living organisms.